Talk: FAIR and Reproducible practices in science¶
Why reproducibility¶
Helps future research: your own and done by others
Facilitates spotting errors and avoids that misconceptions could continue on
Fosters a culture of accountability and trust
Contributes to saving resources. No need to reinvent the wheel
Challenges to reproducibility¶
Complex methodologies that can be difficult to fit into a reproducible environment
Statistical deficiencies (small sample sizes, bias, improper choices)
Societal and cultural issues
Lack of proper training in data management
Publish and perish pressure
Little reward to data sharing
Version Control¶
Management of changes to documents, computer programs and other collections of information
when, who, what
Why should you use it?¶
Benefits¶
transparency
history of changes
backup and restore
recovery from errors
easier collaborative work
reproducibility
Uses¶
You can use version control either alone or collaboratively for:
papers
lectures
documentation
scripts (bash, Python, R or whatever else)
text/CSV/TSV files
Version Control Systems¶
stand-alone tools that record changes to a file or set of files over time
often referred to as VCSs
Basic concept¶
commit
Save files as logical sets of changes and write a good description of why you changed them
What you can do¶
review changes made over time
revert files/the entire project back to a previous state
see who last modified something
find out where and how things went wrong
remove content knowing that you can easily go back
sandboxing
Discussion: should we keep dates or versions in the filenames of a repo?¶
Redundant if we are already tracking changes using the repo
It can make sense when identifying a specific dataset
Quick history¶
~ 40 years since first use
Three main generations. Reference Images
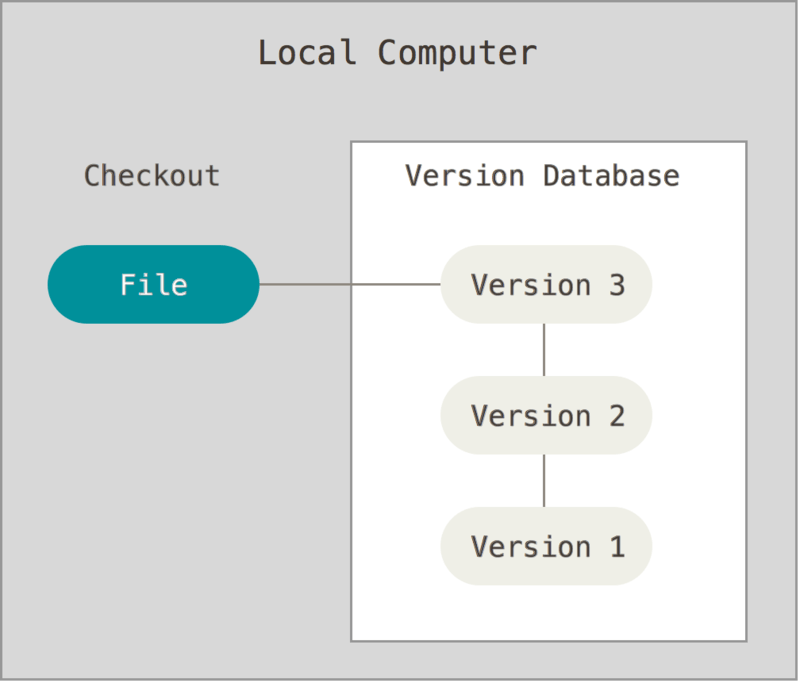
Local¶
e.g. RCS
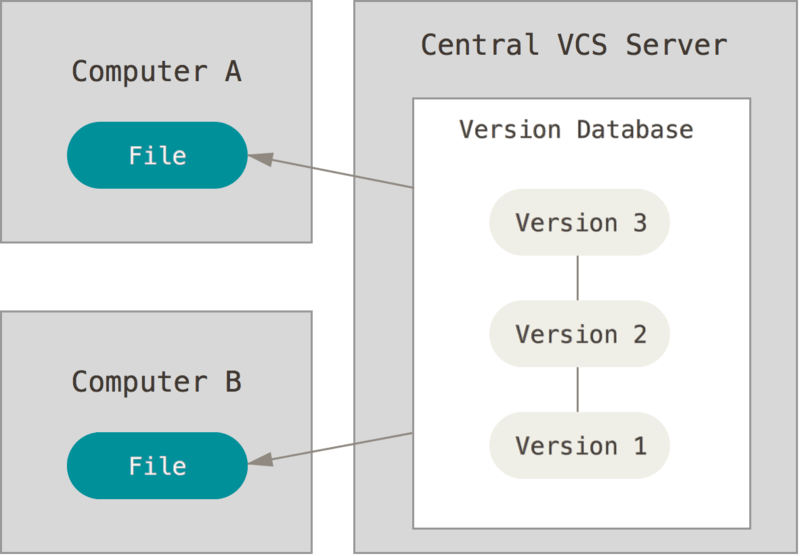
Centralized¶
e.g. SUBVERSION
Distributed¶
e.g. GIT
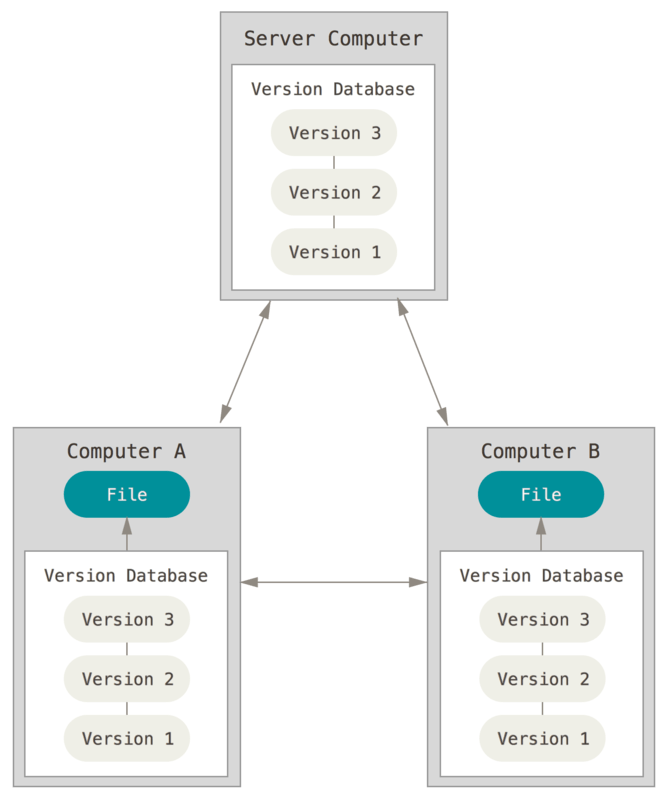
What is Git?¶
An open source, distributed, version control system
It is the most used thanks to its simplicity and GitHub

GitHub is a web-based Git repository hosting service, which offers all the functionalities of Git as well as adding its own features
Git Features¶
nearly every operation is local
integrity, everything is checksummed
generally only adds data
every local repository is a backup
Git hosting services¶
social coding
version control as a service
in-browser editing
additional collaborative features
An alternative to GitHub is GitLab, used mostly in-premise for private projects.
Computational reproducibility¶
Data
Documentation
Software
Software recipes¶
How to wrap all used software and their dependencies
Whenever possible keep specific versions
Help into deployment
Keep as text files
Store in VCS
Containers¶
Software solutions:
Docker (most popular)
Podman (no sysadmin permissions needed)
Singularity/Apptainer (popular in HPC environments)
Docker recipe example, Dockerfile:
FROM debian:bookworm
ARG PERLBREW_ROOT=/usr/local/perl
ARG PERL_VERSION=5.40.0
# Enable perl build options. Example: --build-arg PERL_BUILD="--thread --debug"
ARG PERL_BUILD=--thread
# Base Perl and builddep
RUN set -x; \
apt-get update && apt-get upgrade -y; \
apt-get install -y perl bzip2 zip curl \
build-essential procps
...
Where to find Docker images¶
Docker Hub - Most popular and first registry
Biocontainers - Based on bioconda (more on it later)
Seqera Containers - Creation of containers on demand
Conda¶
Convenient package management system to set up environments
environment.yaml. Recipe for conda environments
conda env create -f myenv.yaml
conda env create myenv
conda env export > myenv.yaml
name: pod5
channels:
- conda-forge
- bioconda
dependencies:
- jannessp::pod5==0.2.4
Tools for installing Conda:
Python¶

requirements.txt
pip install -r requirements.txt
pip freeze > requirements.txt
pandas==1.5.3
ena-upload-cli==0.6.2
Other tools:
R¶

RStudio (.Rproj files)
Renv (renv.lock, along with .Rprofile and renv/activate.R)
Workflows¶
Different steps using different software

RO-Crate - Effort to package research data with metadata.
nf-provproject supports it.Register your workflow or refer to it: WorkflowHub
Example: related to RNA-seq
FAIR¶
More information about the FAIR principles
Findable¶
Example: Dataset available in a public repository with files with clear and descriptive names. Documentation included. Also including tags and keywords to enable its findability.
Counter-example: Dataset not made public. If public, kept in an obscure and non-understandable way without proper tags or documentation.
Accessible¶
Example: Data placed in a well-known format and that can be downloaded in a clear way.
Counter-example: Data encoded in a little known format and with hurdles to download or read.
Interoperable¶
Example: Dataset using terminology, ontology or measurement units that are widely accepted by the community, so it can be compared and integrated with other projects from third parties.
Counter-example: Dataset with project-specific choices that can be hardly mapped to other experiments of the field.
Reusable¶
Example: Dataset provided in a way that contained data can be used easily by third parties and regenerated if needed using available instructions.
Counter-example: Dataset without any documentation, instructions or even context.
RNA-Seq data repositories¶
Public data repositories exist that store data (“raw” and processed) produced by the community from a variety of experiments: microarrays, high-throughput sequencing, high throughput PCR, etc.
It is nowadays required by most journals to make data publicly available upon publication of study in a peer-reviewed journal.
The major repositories for gene expression data:
These repositories are linked to the repositories of NGS raw data (FASTQ files):
Some CLI suggestions¶
CLI tool for downloading data: FASTQ-dl (Container) - Support both SRA and ENA. Can download multiples files from a project
CLI tool for uploading to ENA: ena-upload-cli
First, you upload files to a repository (FTP/Aspera)
Then, upload metadata associated: STUDY, SAMPLE, EXPERIMENT, RUN
Map these 4 concepts to simple TSV files
There are checklists, specific metadata requirements, depending on the sample/experiments that are going to be uploaded: Checklist templates
FAIR in practice for RNA-seq¶
Reference: FAIRification of an RNA-seq dataset
We will use E-MTAB-8316 from ArrayExpress as example.
Findable¶
Example:
Dataset E-MTAB-8316 (from Alivernini et al. 2020, Nature Medicine) has a globally unique and persistent identifier:
E-MTAB-8316or full URLhttps://www.ebi.ac.uk/biostudies/ArrayExpress/studies/E-MTAB-8316Indexed in searchable repositories (ArrayExpress, ENA)
Rich metadata allows discovery via keywords like “macrophage rheumatoid arthritis” combined with “rna-seq of coding rna”
Uses resolution services like identifiers.org for regularized URLs
Counter-example:
RNA-seq data kept only on a lab’s local server without public deposition
FASTQ files named
sample1.fq,sample2.fqwith no associated metadata file
Accessible¶
Example:
Data retrievable via standard HTTP/HTTPS protocol using the persistent identifier
Raw FASTQ files accessible via ENA (European Nucleotide Archive)
Processed data (gene expression matrices) downloadable directly from ArrayExpress
Protocol is open, free, and universally implementable
Counter-example:
Data available only upon request with approval process
FASTQ files in proprietary or obscure sequencing format
Data behind paywall or institutional login requirement
Interoperable¶
Example:
Uses published ontologies: NCBI taxonomy for species (“Homo sapiens”)
Complies with MINSEQE (Minimum Information about a Sequencing Experiment) community standard — 5-star rating
Machine-readable metadata formats:
MAGE-TAB (Sample and Data Relationship Format - SDRF)
ENA SRA XML format
Controlled vocabularies via Annotare submission tool
Counter-example:
Custom gene identifiers not mapped to standard databases (Ensembl, RefSeq)
Proprietary sample annotation system without ontology links
Protocol descriptions using only lab-specific terminology
Reusable¶
Example:
Clear licensing: CC BY 4.0 (Creative Commons Attribution)
Detailed provenance: links to publication, protocol documentation, sample metadata
Complete data availability: raw FASTQ files + processed gene expression matrices
Clear documentation of experimental design, protocols, and variables
Associated with publication DOI: 10.1038/s41591-020-0939-8
Counter-example:
No license specified or restrictive license
Missing protocol details (e.g., “library prep performed as usual”)
Only processed data provided without raw FASTQ files
No link to methods publication