Combining reports
At this point, we can summarize all the work done with the tool MultiQC. MultiQC aggregates outputs from many bioinformatics tools across many samples into a single report by searching a given directory for analysis logs and compiling a HTML report.
MultiQC supports many tools: see the different modules
Let’s create a multiqc_report folder and link all analysis done so far.
cd ~/rnaseq_course/
# create a folder for the multiqc result
mkdir multiqc_report
cd ~/rnaseq_course/multiqc_report
# link QC, trimming and mapping data
ln -s ~/rnaseq_course/quality_control* .
ln -s ~/rnaseq_course/mapping* .
ln -s ~/rnaseq_course/trimming
Then run multiqc on the directory multiqc_report to combine all reports:
cd ~/rnaseq_course/multiqc_report
$RUN multiqc .
We can visualize the final report in the internet browser:
firefox multiqc_report.html
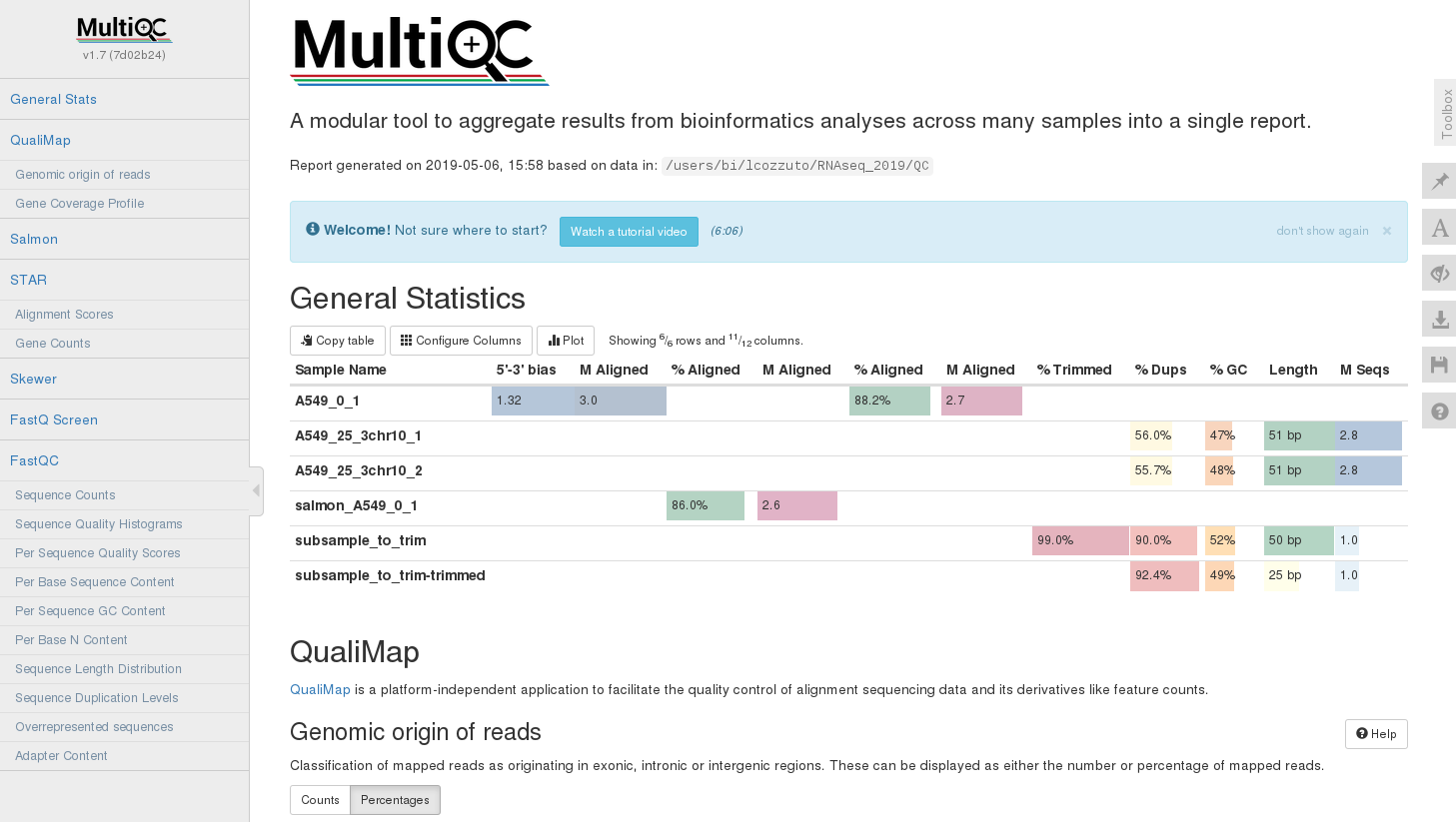
MultiQC is very customizable. You can ignore files or folder:
$RUN multiqc . --ignore trimming --force
# Note that --force must be set in order to allow the overwriting of existing report
Rename output file
$RUN multiqc . --ignore trimming --filename multiqc_notrimming --force
You can export all images
$RUN multiqc . --export --force
# by default, plots are saved in folder "multiqc_plots" in pdf, svg and png format.
ls multiqc_plots/

